DESCRIPTION:
Phase1:
Brain tumor segmentation is to determine the outline of different glioma subregions, which are the enhancing tumor, the tumor core, and the whole tumor. Deep neural networks have achieved great success in this task recently; however, their computational complexity and storage costs hinder their deployment in real-time applications or on resource-constrained devices in clinics. Network pruning, which has attracted many researchers recently, alleviates this limitation by removing trivial parameters of the network. Network pruning is challenging in the medical field because pruning should not degrade the performance of models. As a result, it is vital to choose unimportant parts of networks correctly. This is why I employed the genetic algorithm to identify redundant filters of my U-Net-based network. I considered filter pruning a multiobjective optimization problem in which performance and inference time are optimized simultaneously. Then, I used my compressed network for brain tumor segmentation.
Predicted segmentation masks are often used to estimate patients' survival time. Although several studies have recently achieved good results, they require different feature engineering techniques to extract suitable features, which is difficult and time-consuming. To tackle this problem, I extracted deep features from my compact network and used them for survival prediction.
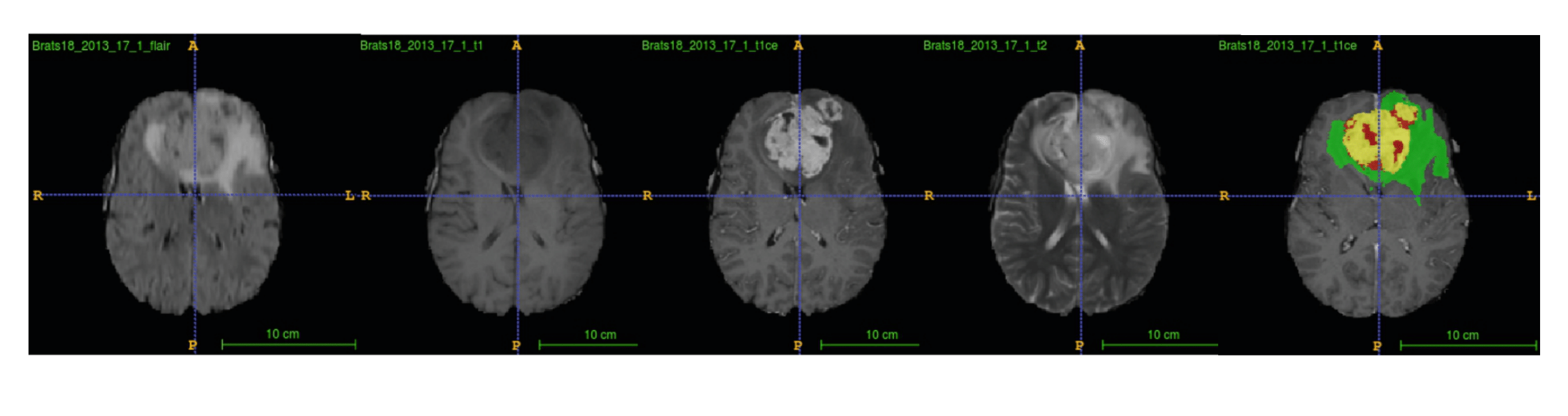
Multimodal MRI with ground truth form BraTS 2018 dataset
The BraTS challenge runs in conjunction with the MICCAI conference each year. I used the BraTS2018 dataset, including 285 MRIs in the training set and 66 MRIs in the validation set. For each case, four MRI sequences(i.e., T1, T1c, T2, and FLAIR volumes) are available in this dataset. The BraTS2018 dataset also provides the survival data of 163 cases in the training set and 53 samples in the validation set. Survival data include patients’ age and resection status.
- Segmentation
- Data preprocessing and augmentation
- Training the segmentation model
- Post-processing
- Network pruning
- Pruning the network using the genetic algorithm
- Overall survival time prediction
- Feature extraction from the pruned network
- Feature selection
- Survival time prediction using machine learning models (SVR, XGBoost, RF, and linear regression)